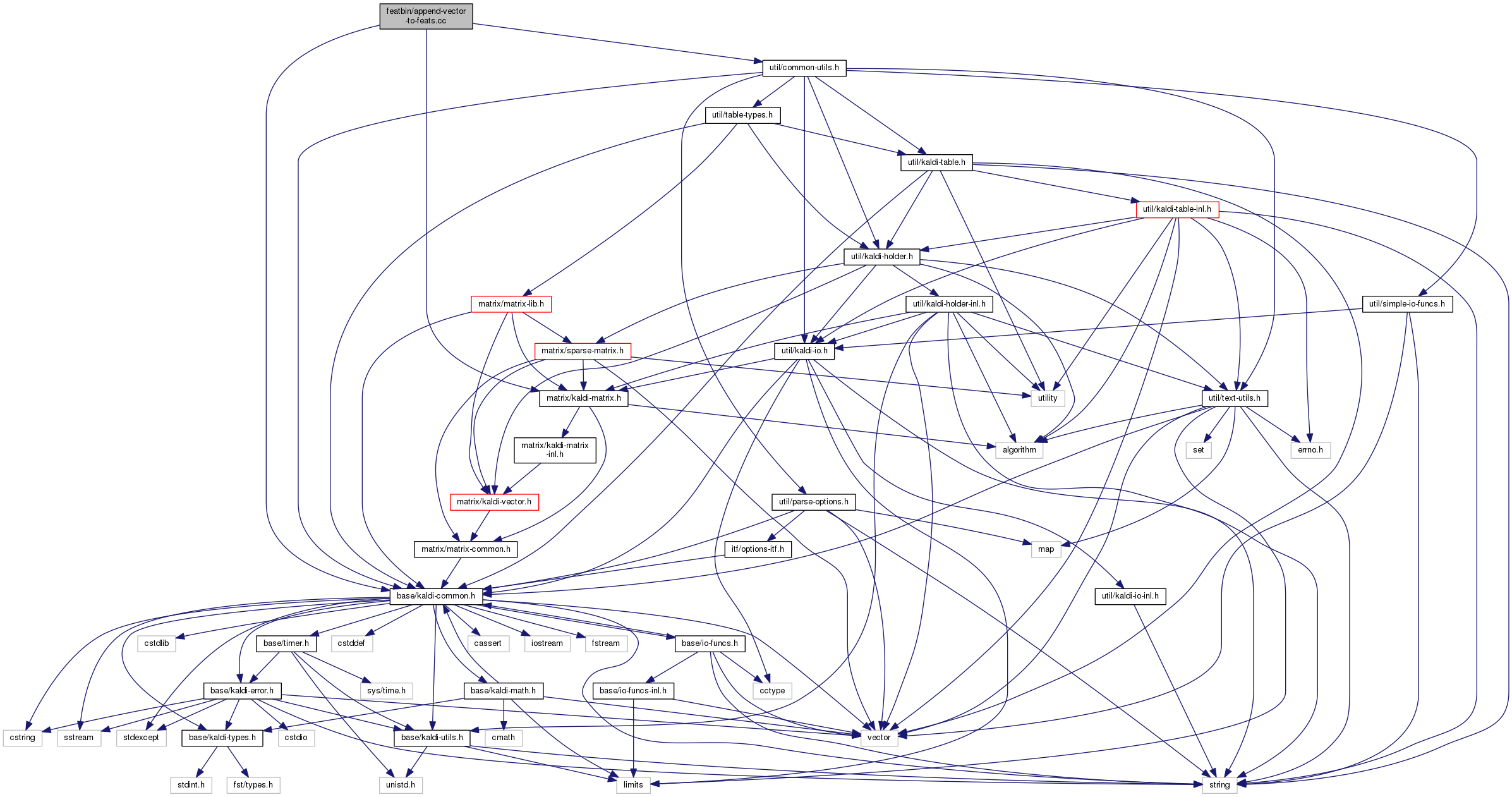
Go to the source code of this file.
Namespaces | |
| kaldi | |
| This code computes Goodness of Pronunciation (GOP) and extracts phone-level pronunciation feature for mispronunciations detection tasks, the reference: | |
Functions | |
| void | AppendVectorToFeats (const Matrix< BaseFloat > &in, const Vector< BaseFloat > &vec, Matrix< BaseFloat > *out) |
| int | main (int argc, char *argv[]) |
| int main | ( | int | argc, |
| char * | argv[] | ||
| ) |
Definition at line 42 of file append-vector-to-feats.cc.
References kaldi::AppendVectorToFeats(), kaldi::ClassifyRspecifier(), SequentialTableReader< Holder >::Done(), ParseOptions::GetArg(), RandomAccessTableReader< Holder >::HasKey(), KALDI_LOG, KALDI_VLOG, KALDI_WARN, SequentialTableReader< Holder >::Key(), kaldi::kNoRspecifier, SequentialTableReader< Holder >::Next(), ParseOptions::NumArgs(), ParseOptions::PrintUsage(), ParseOptions::Read(), kaldi::ReadKaldiObject(), ParseOptions::Register(), RandomAccessTableReader< Holder >::Value(), SequentialTableReader< Holder >::Value(), TableWriter< Holder >::Write(), and kaldi::WriteKaldiObject().