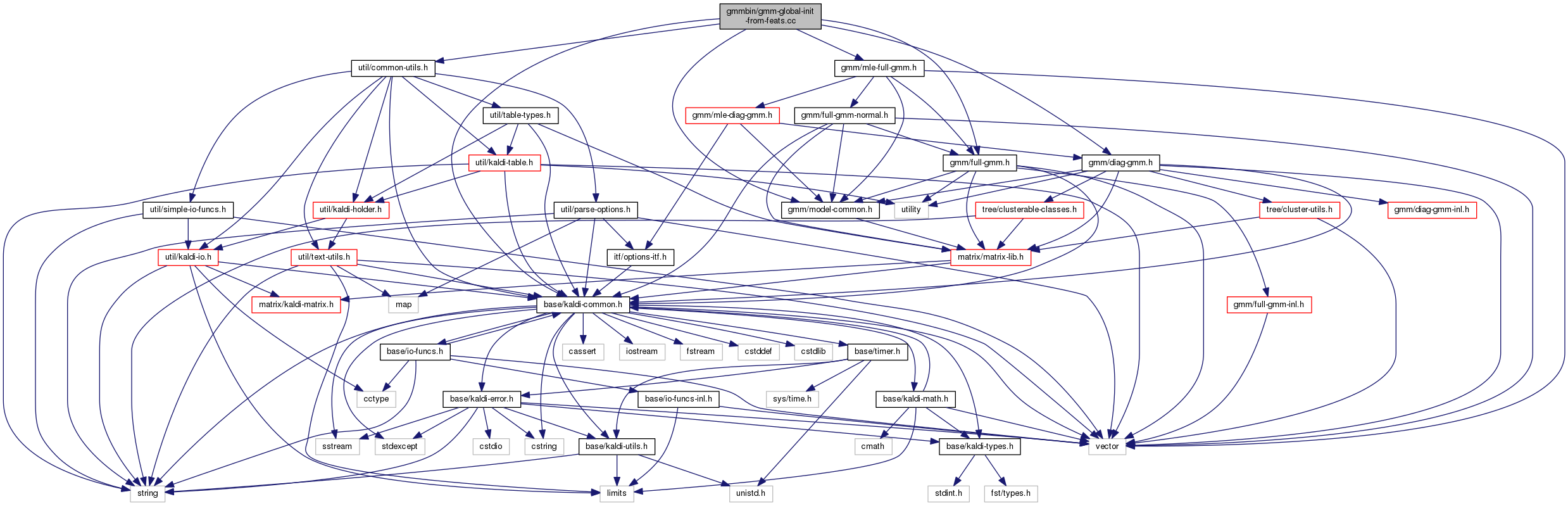
90 using namespace kaldi;
93 "This program initializes a single diagonal GMM and does multiple iterations of\n" 94 "training from features stored in memory.\n" 95 "Usage: gmm-global-init-from-feats [options] <feature-rspecifier> <model-out>\n" 96 "e.g.: gmm-global-init-from-feats scp:train.scp 1.mdl\n";
102 int32 num_gauss = 100;
103 int32 num_gauss_init = 0;
104 int32 num_iters = 50;
105 int32 num_frames = 200000;
106 int32 srand_seed = 0;
107 int32 num_threads = 4;
109 po.
Register(
"binary", &binary,
"Write output in binary mode");
110 po.Register(
"num-gauss", &num_gauss,
"Number of Gaussians in the model");
111 po.Register(
"num-gauss-init", &num_gauss_init,
"Number of Gaussians in " 112 "the model initially (if nonzero and less than num_gauss, " 113 "we'll do mixture splitting)");
114 po.Register(
"num-iters", &num_iters,
"Number of iterations of training");
115 po.Register(
"num-frames", &num_frames,
"Number of feature vectors to store in " 116 "memory and train on (randomly chosen from the input features)");
117 po.Register(
"srand", &srand_seed,
"Seed for random number generator ");
118 po.Register(
"num-threads", &num_threads,
"Number of threads used for " 119 "statistics accumulation");
127 if (po.NumArgs() != 2) {
132 std::string feature_rspecifier = po.GetArg(1),
133 model_wxfilename = po.GetArg(2);
142 int64 num_read = 0, dim = 0;
144 KALDI_LOG <<
"Reading features (will keep " << num_frames <<
" frames.)";
146 for (; !feature_reader.Done(); feature_reader.Next()) {
152 feats.
Resize(num_frames, dim);
153 }
else if (this_feats.
NumCols() != dim) {
154 KALDI_ERR <<
"Features have inconsistent dims " 155 << this_feats.
NumCols() <<
" vs. " << dim
156 <<
" (current utt is) " << feature_reader.Key();
158 if (num_read <= num_frames) {
159 feats.
Row(num_read - 1).CopyFromVec(this_feats.
Row(t));
163 feats.
Row(
RandInt(0, num_frames - 1)).CopyFromVec(this_feats.
Row(t));
169 if (num_read < num_frames) {
170 KALDI_WARN <<
"Number of frames read " << num_read <<
" was less than " 171 <<
"target number " << num_frames <<
", using all we read.";
174 BaseFloat percent = num_frames * 100.0 / num_read;
175 KALDI_LOG <<
"Kept " << num_frames <<
" out of " << num_read
176 <<
" input frames = " << percent <<
"%.";
179 if (num_gauss_init <= 0 || num_gauss_init > num_gauss)
180 num_gauss_init = num_gauss;
182 DiagGmm gmm(num_gauss_init, dim);
184 KALDI_LOG <<
"Initializing GMM means from random frames to " 185 << num_gauss_init <<
" Gaussians.";
190 int32 cur_num_gauss = num_gauss_init,
191 gauss_inc = (num_gauss - num_gauss_init) / (num_iters / 2);
193 for (
int32 iter = 0; iter < num_iters; iter++) {
196 int32 next_num_gauss = std::min(num_gauss, cur_num_gauss + gauss_inc);
197 if (next_num_gauss > gmm.NumGauss()) {
198 KALDI_LOG <<
"Splitting to " << next_num_gauss <<
" Gaussians.";
199 gmm.Split(next_num_gauss, 0.1);
200 cur_num_gauss = next_num_gauss;
205 KALDI_LOG <<
"Wrote model to " << model_wxfilename;
207 }
catch(
const std::exception &e) {
208 std::cerr << e.what();
This code computes Goodness of Pronunciation (GOP) and extracts phone-level pronunciation feature for...
MatrixIndexT NumCols() const
Returns number of columns (or zero for empty matrix).
bool WithProb(BaseFloat prob, struct RandomState *state)
void TrainOneIter(const Matrix< BaseFloat > &feats, const MleDiagGmmOptions &gmm_opts, int32 iter, int32 num_threads, DiagGmm *gmm)
The class ParseOptions is for parsing command-line options; see Parsing command-line options for more...
const SubVector< Real > Row(MatrixIndexT i) const
Return specific row of matrix [const].
void Register(OptionsItf *opts)
A templated class for reading objects sequentially from an archive or script file; see The Table conc...
Configuration variables like variance floor, minimum occupancy, etc.
#define KALDI_ASSERT(cond)
MatrixIndexT NumRows() const
Returns number of rows (or zero for empty matrix).
void InitGmmFromRandomFrames(const Matrix< BaseFloat > &feats, DiagGmm *gmm)
Definition for Gaussian Mixture Model with diagonal covariances.
void WriteKaldiObject(const C &c, const std::string &filename, bool binary)
void Resize(const MatrixIndexT r, const MatrixIndexT c, MatrixResizeType resize_type=kSetZero, MatrixStrideType stride_type=kDefaultStride)
Sets matrix to a specified size (zero is OK as long as both r and c are zero).
int32 RandInt(int32 min_val, int32 max_val, struct RandomState *state)