#include "base/kaldi-common.h"#include "util/common-utils.h"#include "gmm/am-diag-gmm.h"#include "hmm/transition-model.h"#include "fstext/fstext-lib.h"#include "base/timer.h"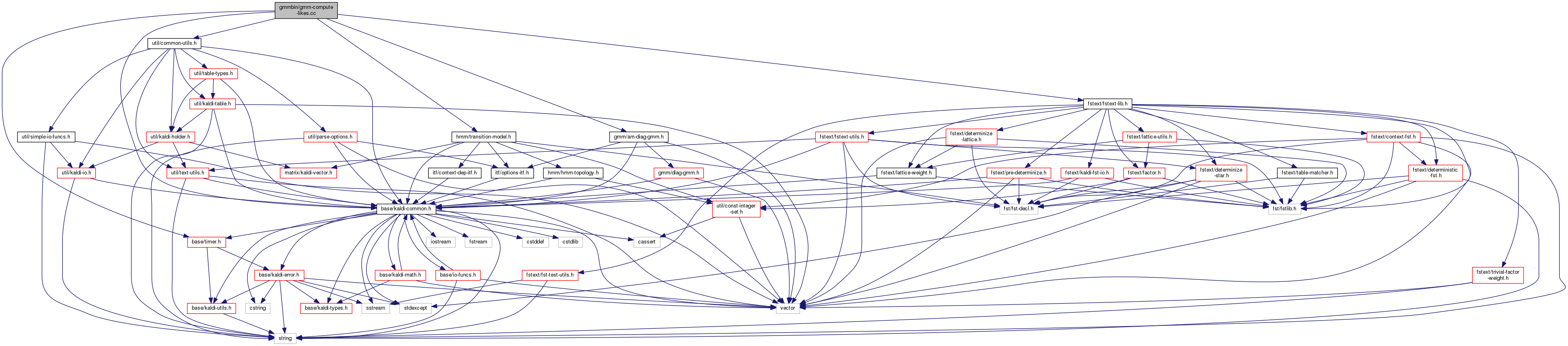
Go to the source code of this file.
Functions | |
| int | main (int argc, char *argv[]) |
| int main | ( | int | argc, |
| char * | argv[] | ||
| ) |
Definition at line 30 of file gmm-compute-likes.cc.
References SequentialTableReader< Holder >::Done(), ParseOptions::GetArg(), rnnlm::i, rnnlm::j, KALDI_LOG, SequentialTableReader< Holder >::Key(), AmDiagGmm::LogLikelihood(), SequentialTableReader< Holder >::Next(), ParseOptions::NumArgs(), AmDiagGmm::NumPdfs(), ParseOptions::PrintUsage(), AmDiagGmm::Read(), ParseOptions::Read(), TransitionModel::Read(), Input::Stream(), SequentialTableReader< Holder >::Value(), and TableWriter< Holder >::Write().